A Ghost Painting Coming to Life in the Studio of the Painter Okyō, from the series Yoshitoshi ryakuga (Sketches by Yoshitoshi). 1882 print by Tsukioka Yoshitoshi. Vanderbilt University Fine Arts Gallery 1992.083 via Wikimedia Commons. Wikidata item Q102961245
For several years, I've been working with the Vanderbilt Fine Arts Gallery staff to create and improve Wikidata items for the approximately 7000 works in the Gallery collection through the WikiProject Vanderbilt Fine Arts Gallery. In the past year, I've been focused on creating a Python script to streamline the process of uploading images of Public Domain works in the collection to Wikimedia Commons, where they will be freely available for use. I've just completed work on that script, which I've called CommonsTool, and have used it to upload over 1300 images (covering about 20% of the collection and most of the Public Domain artworks that have been imaged).
In this post, I'll begin by describing some of the issues I dealt with and how they resulted in features of the script. I will conclude by outlining briefly how the script works.
The script is freely available for use and there are detailed instructions on GitHub for configuring and using it. Although it's designed to be usable in contexts other than the Vanderbilt Gallery, it hasn't been tested thoroughly in those circumstances. So if you try using it, I'd like to hear about your experience.
Wikidata, Commons, and structured data
If you have ever worked with editing metadata about art-related media in Wikimedia Commons, you are probably familiar with the various templates used to describe the metadata on the file page using Wiki syntax. Here's an example:
=={{int:filedesc}}==
{{Artwork
|artist = {{ Creator | Wikidata = Q3695975 | Option = {{{1|}}} }}
|title = {{en|'''Lake George'''.}}
|description = {{en|1=Lake George, painting by David Johnson}}
|depicted people =
|depicted place =
|date =
|medium = {{technique|oil|canvas}}
|dimensions = {{Size|in|24.5|19.5}}
|institution = {{Institution:Vanderbilt University Fine Arts Gallery}}
|references = {{cite web |title=Lake George |url=https://library.artstor.org/#/asset/26754443 |accessdate=30 November 2020}}
|source = Vanderbilt University Fine Arts Gallery
|other_fields =
}}
=={{int:license-header}}==
{{PD-Art|PD-old-100-expired}}
[[Category:Vanderbilt University Fine Arts Gallery]]
These templates are complicated to create and difficult to edit by automated means. In recognition of this, the Commons community has been moving towards storing metadata about the media files as structured data ("Structured Data on Commons", SDC). When media files depict artwork, the preference is to describe the artwork metadata in Wikidata rather than as wikitext on the Commons file page (as shown in the example above).
In July, Sandra Fauconnier gave a presentation at an ARLIS/NA (Art Libraries Society of North America) Wikidata group meeting that was extremely helpful for improving my understanding of the best practices for expressing metadata about visual artworks in Wikimedia Commons. She provided a link to a very useful reference page (still under construction as of September 2022) to which I referred while working on my script.
The CommonsTool script has been designed around two key features for simplifying management of the media and artwork metadata. The first is two very simple wikitexts: one for two-dimensional artwork and another for three-dimensional artwork. The 2D wikitext looks like this:
=={{int:filedesc}}==
{{Artwork
|source = Vanderbilt University
}}
=={{int:license-header}}==
{{PD-Art|PD-old-100-expired}}
[[Category:Vanderbilt University Fine Arts Gallery]]
and the 3D wikitext looks like this:
=={{int:filedesc}}==
{{Art Photo
|artwork license = {{PD-old-100-expired}}
|photo license = {{Cc-by-4.0 |1=photo © [https://www.vanderbilt.edu/ Vanderbilt University] / [https://www.library.vanderbilt.edu/gallery/ Fine Arts Gallery] / [https://creativecommons.org/licenses/by/4.0/ CC BY 4.0]}}
}}
[[Category:Vanderbilt University Fine Arts Gallery]]
By comparison with the wikitext in the first example, this is clearly much simpler, but also has the advantage that there is very little metadata in the wikitext itself that might need to be updated.
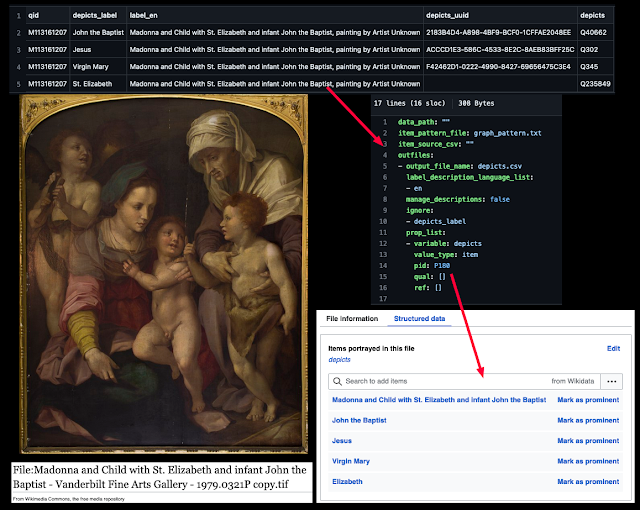
The second key feature involves using SDC to link the media file to the Wikidata item for the artwork. Here's an example for the work shown at the top of this post:
In order for this strategy to work, for all artwork images the depicts (P180) and main subject (P921) values must be set to the artwork's Wikidata item (in this case Q102961245). Two dimensional artwork images should also have a "digital representation of" (P6243) value with the artwork's Wikidata item. When these claims are created, the Wikidata metadata will "magically" populate the file information summary without entering it into a wikitext template.
The great advantage here is that when metadata are updated on Wikidata, they automatically are updated in Commons as well.
Copyright and licensing issues
One of the complicating issues that had slowed me down in developing the script was to figure out how to handle copyright and licensing issues. The images we are uploading depict old artwork that is out of copyright, but what about copyright of the images of the artwork? The Wikimedia Foundation takes the position that faithful photographic reproductions of old two-dimensional artwork lack originality and are therefore not subject to copyright. However, images of three-dimensional works can involve creativity, so those images must be usable under an open license acceptable for Commons uploads.
Wikitext tags
Unlike other metadata properties about a media item, the copyright and licensing details cannot (as of September 2022) be expressed only in SDC. They must be explicitly included in the file page's wikitext.
As shown in the example above, I used the license tags
{{PD-Art|PD-old-100-expired}}
for 2D artwork. The PD-Art tag asserts that the image is not copyrightable for the reason given above and PD-old-100-expired asserts that the artwork is not under copyright because it is old. When these tags are used together, they are rendered on the file page like this:
The example above for 3D artworks uses separate license tags for the artwork and the photo. The artwork license is
PD-old-100-expired as before, and the photo license I used was
{{Cc-by-4.0 |1=photo ©
[https://www.vanderbilt.edu/ Vanderbilt University] /
[https://www.library.vanderbilt.edu/gallery/ Fine Arts Gallery] /
[https://creativecommons.org/licenses/by/4.0/ CC BY 4.0]}}
There are a number of possible licenses that can be used for both the photo and artwork and they can be set in the CommonsTool configuration file. Since the CC BY license requires attribution, I used the explicit credit line feature to make clear that it's the photo (not the artwork) that's under copyright and to provide links to Vanderbilt University (the copyright holder) and the Fine Arts Gallery. Here's how these tags are rendered on the file page of an image of a 3D artwork:
Using the format
{{Art Photo
|artwork license = {{artLicenseTag}}
|photo license = {{photoLicenseTag}}
}}
in the wikitext is great because it creates separate boxes that clarify that the permissions for the artwork are distinct from the permissions for the photo of the artwork.
Structured data about licensing
As noted previously, it's required to include copyright and licensing information in the page wikitext. However, file pages must also have certain structured data claims related to the file creator, copyright, and licensing or they will be flagged.
In the case of 2D images where the PD-Art tag was used, there should be a "digital representation of" (P6243) claim where the value is the Q ID of the Wikidata item depicted in the media file.
In the case of 3D images, they should not have a P6243 claim, but should have values for copyright status (P6216) and copyright license (P275). If under copyright, they should also have values for creator (P170, i.e. photographer) and inception (P571) date so that it can be determined to whom attribution should be given and when the copyright may expire. Keep in mind that for artwork SDC metadata is generally about the media file and not the depicted thing. So similar information about the depicted artwork would be expressed in the Wikidata item about the artwork, not in SDC.
Although not required when the PD-Art tag is used, it's a good idea to include the creator (photographer) and inception date of the image in the SDC metadata for 2D works. It's not yet clear to me whether a copyright status value should be provided. I suppose so, but if it's directly asserted in the SDC that the work is in the Public Domain, you are supposed to use a qualifier to indicate the reason, and I'm not sure what value would be used for that. I haven't seen any examples illustrating how to do that, so for now, I've omitted it.
To see examples of how this looks in practice see this example for 2D and this example for 3D. After the page loads, click on the Structured Data tab below the image.
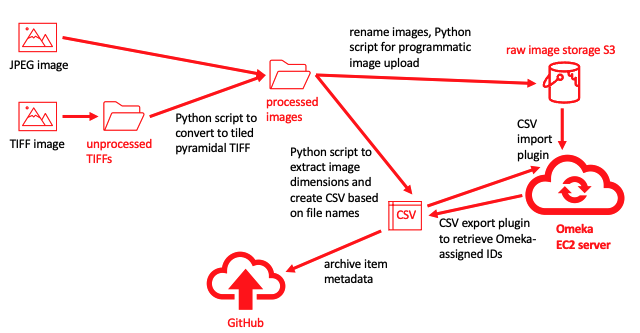
What the script does: the Commons upload
The Commons upload takes place in three stages.
First, CommonsTool acquires necessary information about the artwork and the image from CSV tables. One key piece of information is what image or images to be uploaded to Commons are associated with a particular artwork (represented by a single Wikidata item). The main link from Commons to Wikidata is made using a depicts (P180) claim in the SDC and the link from Wikidata to Commons is made using an image (P18) claim.
Miriam by Anselm Feuerbach. Public Domain via Wikimedia Commons
It is important to know whether there are more than one image associated with the artwork. In the source CSV data about images, the image to be linked from Wikidata is designated as "primary" and additional images are designated as "secondary".
Both primary and secondary images will be linked from Commons to Wikidata using a depicts (P180) claim, but it's probably best for only the primary image to be linked from Wikidata using an image (P18) claim. Here is an example of a primary image page in Commons and here is an example of a secondary image page in Commons. Notice that the Wikidata page for the artwork only displays the primary image.
The CommonsTool script also constructs a descriptive Commons filename for the image using the Wikidata label, any sub-label particular to one of multiple images, the institution name, and the unique local filename. There are a number of characters that aren't allowed, so CommonsTool tries to find them and replace them with valid characters.
The script also performs a number of optional screens based on copyright status and file size. It can skip images deemed to be too small and will also skip images whose file size exceeds the API limit of 100 Mb. (See the configuration file for more details.)
The second stage is to upload the media file and the file page wikitext via the Commons API. Commons guidelines state that the rate of file upload should not be greater than one upload per 5 seconds, so the script introduces a delay of necessary to avoid exceeding this rate. If successful, the script moves on to the third stage and if not, it logs an error and moves to the next media item.
In the third stage, SDC claims are written to the API in a manner similar to how claims are written to Wikidata. The claims upload function respects the maxlag errors from the server and delays the upload if the server is lagged due to high usage (although this rarely seems to happen). If the SDC upload fails, it logs an error, but the script continues in order to record the results of the media upload in the existing uploads CSV file.
The links from the Commons image(s) to Wikidata are made using SDC statements, which results in a hyperlink in the file summary (the tiny Wikidata flag). However, the link in the other direction doesn't get made by CommonsTool.
The CSV file where existing uploads are recorded contains an image_name column and the primary values for "primary" images in that column can be used as values for the image (P18) property on the corresponding Wikidata artwork item page. After creating that claim, the primary image will be displayed on the artwork's Wikidata page:
Making this link manually can be tedious, so there is a script that will automatically transfer these values into the appropriate column of a CSV file that is set up to be used by the VanderBot script to upload data to Wikidata. In production, I have a shell script that runs CommonsTool, then the transfer script, followed by VanderBot. Once that shell script has finished running, the image claim will be present on the appropriate Wikidata page.
International Image Interoperability Framework (IIIF) functions
One of our goals at the Vanderbilt Libraries (of which the Fine Arts Gallery is part) is to develop the infrastructure to support serving images using the International Image Interoperability Framework (IIIF). To that end, we've set up a Cantaloupe image server on Amazon Web Services (AWS). The setup details are way beyond the scope of this web post, but now that we have this capability, we want to make the images that we've uploaded to Commons also be available as zoomable high-resolution images via our IIIF server.
For that reason, the CommonsTool script also has the capacity to upload images to the IIIF server storage (an AWS bucket) and to generate manifests that can be used to view those images. The IIIF functionalities are independent of the Commons upload capabilities -- either can be turned on or off. However, for my workflow, I do the IIIF functions immediately after the Commons upload so that I can use the results in Wikidata as I'll describe later.
Source images
One of the early things that I learned when experimenting with the server is that you don't want to upload large, raw TIFF files (i.e. greater than 10 MB). When a IIIF viewer tries to display such a file, it has to load the whole file, even if the screen area is much smaller that the entire TIFF would be if displayed at full resolution. This takes an incredibly long time, making viewing of the files very annoying. The solution to this is to convert the TIFF files into tiled pyramidal TIFFs.
When I view one of these files using Preview on my Mac, it becomes apparent why they are called "pyramidal". The TIFF file doesn't contain a single image. Rather, it contains a series of images that are increasingly small. If I click on the largest of the images (number 1), I see this:
and if I click on a smaller version (number 3), I see this:
If you think of the images as being stacked with the smaller ones on top of the larger ones, you can envision a pyramid.
When a client application requests an image from the IIIF server, the server looks through the images in the pyramid to find the smallest one that will fill up the viewer and sends that. If the viewer zooms in on the image, requiring greater resolution, the server will not send all of the next larger image. Since the images in the stack are tiled, it will only send the particular tiles from the larger, higher resolution image that will actually be seen in the viewer. The end result is that the tiled pyramidal TIFFs load much faster because the IIIF server is smart and doesn't send any more information than is necessary to display what the user wants to see.
The problem that I faced was how to automate the process of generating a large number of these tiled pyramidal TIFFs. After thrashing with various Python libraries, I finally ended up using the command line tool ImageMagick and calling it from a Python script using the os.system() function. The script I used is available on GitHub.
Because the Fine Arts Gallery has been working on imaging their collection for over 20 years, the source images that I'm using are in a variety of formats and sizes (hence the optional size screening criteria in the script to filter out images that have too low resolution). The newer images are high resolution TIFFs, but many of the older images are JPEGs or PNGs. So one task of the IIIF server upload part of the CommonsTool script is to sort out whether to pull the files from the directory where the pyramidal TIFFs are stored, or the directory where the original images are stored.
Once the location of the correct images are identified, the script uses the boto3 module (the AWS software development kit or SDK), to initiate the upload to the S3 bucket as part of the Python script. I won't go into the details of setting up and using credentials as that is described well in the AWS documentation.
Once the file is uploaded, it can be directly accessed using a URL constructed according to the IIIF Image API standard. Here's a URL you can play with:
https://iiif.library.vanderbilt.edu/iiif/3/gallery%2F1992%2F1992.083.tif/full/!400,400/0/default.jpg
If you adjust the URL (for example replacing the 400s with different numbers) according to the API 2.0 URL patterns, you can make the image display at different sizes directly in the browser.
IIIF manifests
The real reason for making images available through a IIIF server is to display them in a viewer application. One such application is Mirador. A IIIF viewer uses a manifest to understand how the image or set of images should be displayed. CommonsTool generates very simple IIIF manifests that display each image in a separate canvas, along with basic metadata about the artwork. To see what the manifest looks like for the image at the top of this post, go to this link.
IIIF manifests are written in machine-readable Javascript Object Notation (JSON), so they are not intended to be understood by humans. However, when the manifest is consumed by a viewer application, a human can use controls such as pan, zoom, and buttons to manipulate the image or to move to another canvas that displays a different image. The Mirador project provides an online IIIF viewer that can be used to view images described by a manifest. This link will display the manifest from above in the Mirador online viewer.
One nice thing about providing a IIIF manifest is that it allows multiple images of the same work to be viewed in the same viewer. For example, there might be multiple pages of a book, or the front and back sides of a sculpture. I'm still learning about constructing IIIF manifests, so I haven't done anything fancy yet with respect to generating IIIF manifests in the CommonsTool script. However, the script does generate a single manifest describing all of the images depicting the same artwork. The image designated as "primary" is shown in the initial view and any other images designated as "secondary" are shown in other canvases that can be selected using the viewer display options or be viewed sequentially using the buttons at the bottom of the viewer. Here is an example showing how the manifest for the primary and secondary images in an earlier example put the front and back images of a manuscript page in the same viewer window.
IIIF in Wikidata
Wikidata has a property "IIIF manifest" (P6108) that allows an item to be linked to a IIIF manifest that displays depictions of that item. The file where existing uploads are recorded includes a iiif_manifest column that contains the manifest URLs for the works depicted by the images.
Those values can be used to create IIIF manifest (P6108) claims for an item in Wikidata:
Because doing this manually would be tedious, the iiif_manifest values can be automatically transferred to a VanderBot-compatable CSV file using the same transfer script used to transfer the image_name.
In itself, adding a IIIF manifest claim isn't very exciting. However, Wikidata supports a user script that will display an embedded Mirador viewer anytime an item has a value for P6108. (For details on how to install that script, see this post.) With the viewer enabled, opening a Wikidata page for a Fine Arts Gallery item with images will display the viewer at the top of the page and a user can zoom in or use the buttons at the bottom to move to another image of the same artwork.
This is really nice because if only the primary image is linked using the image property, users would not necessarily know that there are other images of the object in Commons. But with the embedded viewer, the user can flip through all of the images of the item that are in Commons using the display features of the viewer, such as thumbnails.
Using the script
Although I wrote this script primarily to serve my own purposes, I tried to make it clean and customizable enough that someone with moderate computer skills should also be able to use it. The only installation requirements are Python and several modules that aren't included in the standard library. It should not generally be necessary to modify the script to use it -- most customizing should be possible by changing the configuration file.
If the script is only used to write files to Commons, it's operation is pretty straightforward. If you want to combine uploading image files to Commons with writing the image_names and iiif_manifest values to Wikidata, it's more complicated. You need to get the transfer_to_vanderbot.py script working and then learn how to operate VanderBot. There are details instructions, videos, etc. to do that on the VanderBot landing page.
What's next?
There are still a few more Fine Arts Gallery images that I need to upload after doing some file conversions, checking out some copyright statuses, and wranging some data for multiple files that depict the same work. However, I'm quite excited about developing better IIIF manifests that will make it possible to view related works in the same viewer. Having so many images in Commons now also makes it possible to see the real breadth of the collection by viewing the Listeria visualizations on the tabs of the WikiProject Vanderbilt Fine Arts Gallery website. I hope soon to create more fun SPARQL-based visualizations to add to those already on the website landing page.













_-_Vanderbilt_Fine_Arts_Gallery_-_1992.083.tif/lossy-page1-782px-thumbnail.tif.jpg)














